What is LabID?
LabID (Lab Integrated Data) is an open-source web-based platform for research data management in life science institutes, featuring sample and dataset management, an inventory management system and an electronic lab notebook.
LabID enables the recording of extensive experimental information about the provenance of data (samples, reagents, instruments, protocols, and assay parameters), and is designed to help individual scientists, research groups, and core facilities better manage, annotate, and share their research according to the FAIR principles.
LabID also features an electronic lab notebook, and a “workflow integration” to keep track of the execution of workflows such as Galaxy or Nextflow. Workflows can be imported from various sources: Git-based platforms such as GitHub, GitLab, and Bitbucket, Galaxy instances, WorkflowHub or any platform that can provide workflows using the Workflow RO-Crate specification. Workflows can also be registered in LabID by simple drag-and-drop of local files. Similarly, LabID supports importing and exporting workflow runs using the interoperable Workflow Run RO-Crate specification, which allows capturing both the data used and generated by the workflow, and metadata about the workflow execution (version of the workflow, parameter values, etc.). Such Research Object Crate (RO-Crate) can be deposited to repositories such as Zenodo or RoHub.
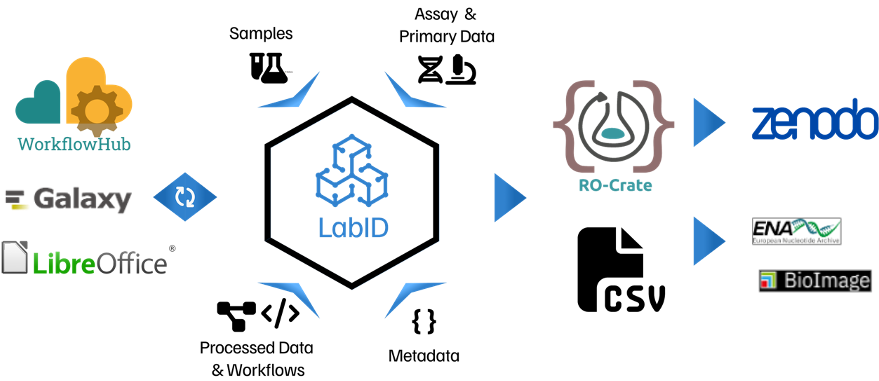
Overview of LabID functionalities
Key Features
LabID is powered by a database that allows recording and interconnecting laboratory entities involved in life-science research (e.g. samples, reagents, instruments, and protocols). The result is a comprehensive knowledge graph capturing relationships between experimental components, enabling researchers to trace data provenance and share the full context of their research.
- Sample Management: Track and organise biological samples with complete provenance information
- Dataset Management: Register and annotate datasets with metadata and experimental parameters
- Electronic Lab Notebook: Document experiments and observations
- Inventory System: Manage reagents, instruments, and other lab resources
- Workflow Integration: Keep track of workflow versions and workflow executions for platforms like Galaxy or Nextflow, and custom scripts. Import and export workflows from/to platforms like WorkflowHub and Git-based platforms such as GitHub, GitLab, and Bitbucket.
You can find a video with an overview of LabID’s features here. The video dates back to the time the software was called “Stocks”. The features are the same, though, and the interface has only changed slightly since. The LabID workflow integration is also summarised in the following video.
Which tasks can be solved with LabID?
- Referencing instruments, reagents, and specimens available in an institute or research team
- Documenting assays performed in the lab, associating them to the instrument, reagent and specimen used (e.g imaging of some tissue on a microscope)
- Storing and archiving of raw and processed data (e.g archiving a copy to an S3 bucket)
- Adding metadata to any entity using custom or controlled vocabulary
- Sharing data, protocols, and assays with internal colleagues
- Document the execution of a script or workflow, together with the associated data and parameters. Export the resulting “workflow run” as a Workflow Run RO-Crate.
How to access LabID?
LabID is typically installed at the institute or research group level on a Unix server. The LabID user interface is accessible via a web browser, on a client machine that can access the server.
A public demo server is available at https://labid-demo.embl.de/ so you can get a feeling of the user interface. Most trainings from the user-documentation can also be followed along with this demo server.
What technology is it built with?
Under the hood, LabID is a client/server solution, similar in its architecture to e.g OMERO. It is made of several components :
- the server or “backend”, written in Python and using the Django framework
- a postgres relational database, used to store reference to data and metadata
- the user interface or “frontend”, written in Vue.js
- a Python library and command line interface, to automate e.g. data-registration tasks
Links
Funding
LabID is developed and maintained by the MODIS team at the EMBL Heidelberg.
The workflow integration of LabID was supported through the Open Science Clusters’ Action for Research and Society (OSCARS) European project under grant agreement Nº101129751. See LabID PROV.
Related pages
More information
Training
Skip tool tableTools and resources on this page
| Tool or resource | Description | Related pages | Registry |
|---|---|---|---|
| Bitbucket | Git-based code hosting and collaboration tool, built for teams. | Data organisation | Standards/Databases |
| Galaxy | Open, web-based platform for data intensive biomedical research. Whether on the free public server or your own instance, you can perform, reproduce, and share complete analyses.
|
Marine Metagenomics Single-cell sequencing Virology Data analysis Data provenance Data storage | Tool info Training |
| Git | Distributed version control system designed to handle everything from small to very large projects | Single-cell sequencing Data organisation | Training |
| GitHub | Versioning system, used for sharing code, as well as for sharing of small data | Biodiversity Machine learning Creating a data-flow d... Data organisation Documentation and meta... | Standards/Databases Training |
| GitLab | GitLab is an open source end-to-end software development platform with built-in version control, issue tracking, code review, CI/CD, and more. Self-host GitLab on your own servers, in a container, or on a cloud provider. | Data organisation | Standards/Databases Training |
| LabID (Lab Integrated Data) | LabID is a data management solution for life-science institutes, featuring sample and dataset management, an inventory management system and an electronic lab notebook. | Data organisation Data provenance Documentation and meta... | Tool info |
| Nextflow | Nextflow is a framework for data analysis workflow execution | Data analysis | Tool info Training |
| OMERO | OMERO is an open-source client-server platform for managing, visualizing and analyzing microscopy images and associated metadata | Galaxy OMERO Bioimaging data Data organisation | Tool info Training |
| Research Object Crate (RO-Crate) | RO-Crate is a lightweight approach to packaging research data with their metadata, using schema.org. An RO-Crate is a structured archive of all the items that contributed to the research outcome, including their identifiers, provenance, relations and annotations. | Galaxy Microbial biotechnology Plant sciences Data provenance | Standards/Databases Training |
| RoHub | RoHub is a solution for the storage, life-cycle management and preservation of scientific investigations, campaigns, and operational processes via research objects. | Standards/Databases | |
| WorkflowHub | WorkflowHub is a registry for describing, sharing and publishing scientific computational workflows. | Galaxy Biodiversity Data analysis Data provenance | Tool info Standards/Databases Training |
| Zenodo | Generalist research data repository built and developed by OpenAIRE and CERN | FAIRtracks Plant Phenomics Bioimaging data Biomolecular simulatio... Enzymology and biocata... Plant sciences Single-cell sequencing Data publication Identifiers | Standards/Databases Training |
